With the growing risk of the fast-paced spread of COVID pandemic across the globe, there is an urgent need for potential approaches to break the chain, if not all-out, to find a cure. Currently, there is significant research and literature for this uncertain situation which is majorly related to previous epidemics spread. It might not be totally specific for our current pandemic situation but still can be valuable. These works of literature and research can provide information and approaches which can be used to improve the policy measures to fight this pandemic.
Thus, researchers — Prathamesh P. Karmalkar, a principal data scientist at Merck Group; Rohit Rangarajan, an NLP Expert at Merck Group; Bharat Hegde, master thesis project at Merck KGaA; Dr Harsha Gurulingappa, text analytics product owner at Merck KGaA; and Jerry Megaro, global head of advanced analytics at EMD Millipore Corporation — have developed an NLP-based search engine leveraging these research, ideas and data available to find accurate COVID actionable insights for bringing out medical innovation. With this solution, the researchers are aiming to help the community to find the right information using the methods of deep learning search.
Also Read: How This AI Model Can Get Students Back In School Premise In The Post-COVID World
NLP & Deep Learning-Enabled Engine To The Rescue
An in-house solution developed as a part of Merck Group’s research and development capabilities and activities has been designed to retrieve necessary COVID related information from the massive dataset of literature articles. Renowned as a multinational pharmaceutical company, Merck Group has always been prominently into smart technologies.
According to researchers, the dataset is exponentially growing every day and therefore becomes a hurdle to train the AI/ML system every day on the new evolving data. “Also, it becomes tedious for researchers and innovators to tackle the language nuances by manually annotating the articles,” said Prathamesh, one of the researchers from Merck Group. And thus, as a part of the ‘Text Retrieval Challenge,’ the NLP researchers built an NLP model for extracting information from the enormous COVID-19 dataset of literature articles available.
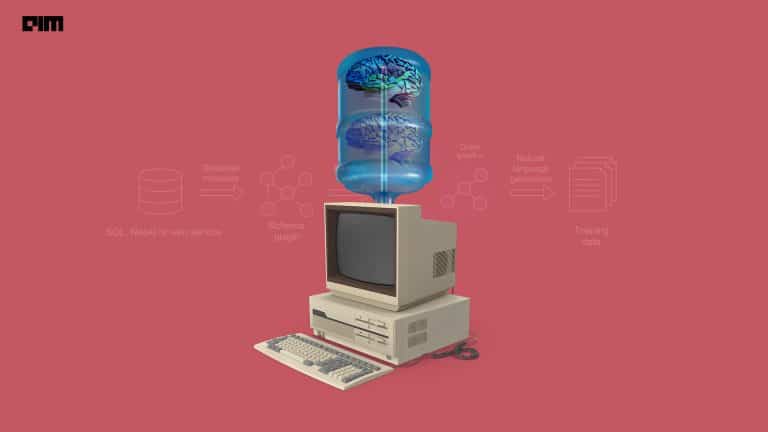
The researchers developed an NLP and deep learning-enabled engine which can accept natural language and free text dynamic queries to acquire relevant information from the offline repositories of 186,000 articles from PubMed Central, WHO, bioRxiv, and medRxiv corpora.
Prathamesh further explained — once the query has been put into the search engine, the algorithm works its way up to highlight the specific sentences and sections from the article where the answer of the input query can be found. Alongside, the model also computes the confidence score associated with every hit to determine the score for each hit corresponding to the given input query.
The researchers also came up with a semantic search-based information retrieval system using Facebook AI Semantic Search (FAISS) and Universal Sentence Encoder (USE).
The model developed by the researchers has therefore been trained and optimised for sentence-level information, where input is of variable-length of an English sentence, and the output is a 512-dimensional vector. “We started with Sentence Transformer for the embeddings, but it had 768-dimension representation, and the final index occupied approximately 20GB. Also, the time taken to generate the embeddings for 7 million sentences was approximately 3.5 hours with K80 GPU on AWS SageMaker,” explained Prathamesh. However, the USE approach has 512-dimensional vector embeddings, thus occupying less space of only 16GB, and took only 20 minutes to compute all the embeddings.
Further, the researcher leveraged the Facebook developed FAISS tool for indexing their sentence embeddings. The researchers were required to reduce their storage space to almost 16 GB, and for this, they approached three techniques — product quantisation, scalar quantisation and IndexIVFFlat.
Firstly, for product quantisation techniques, the researchers divided 512 dimension vectors into eight parts of 64 dimensions each, with which the clustering was done to get 64 centroids. Each vector chunk has been assigned to the closest cluster centroid, which made the vector eight-dimensional. Secondly, the scalar quantisation technique was used to reduce it to 32-bit float into a six-bit vector representation.
And, thirdly, the researchers used IndexIVFFlat, which trained the index where clustering is performed on all vectors to form ‘k’ clusters. Explaining further, Prathamesh stated, “Although the search time was reduced using this approach, we needed to be careful here as there could have been a potential trade-off with accuracy.”
According to the researchers, a regular search operation takes 1.8 secs in ml.m5.4xlarge SageMaker, however with the elastic search EC2 instance, it took only six seconds. And therefore, Elasticsearch (ES) 7.6 was investigated as an alternative to FIASS.
How it works across all documents.
Additionally, the distributed indexing, as well as enterprise features such as security, fail-safe, encryption, made the solution even more suitable to operate within an industrialised environment. The researchers applied text processing techniques to clean the noise and to get a better quality text from the search engine.
What Are The Benefits & Future Plan
According to researchers, with the above pointers in hand, it is believed that using Elasticsearch and FIASS together can bring benefits in solving search problems on the miscellaneous biomedical literature articles.
This is because — while FAISS builds a semantic/vector-space index to identify K closest vectors to the one representing a query, Elasticsearch builds inverted-index in a tree-like fashion to analyse and sync a keyword to the documents list containing the keyword. Also, FAISS not only provides optimisation algorithms to speed up the search but also supports GPU for indexing and search. Having said that, while FAISS only allows searching on vectors, ES will enable the search to happen on the text as well as on embeddings/vectors.
The researchers believe that the solution can prove to be immensely beneficial for organisations, especially in the healthcare industry, to search and find pieces of evidence for any COVID-19 related questions. As part of the further phases of this solution, the researchers are working towards adding the functionality of QnA system that would fetch exact answers to questions, instead of longer sections where the user must find the answer. “We hope to improve our Q&A system using reinforcement learning techniques to enhance the retrieval process of the engine,” concluded Prathamesh.