Researchers from Google’s DeepMind and Alphabet’s Calico have collaborated to introduce a neural network architecture — Enformer. It’s a transformers-based model with the ability to predict gene expression from DNA sequences with greater accuracy. Simply put, Gene expression is nothing but the process in which DNA directs the synthesis of proteins that underpin every biological process in the human body. These developments outline the ability of artificial intelligence to offer unique benefits for human health and accelerate scientific progress.
Published today in @naturemethods together with colleagues from @calico: Enformer – a transformer model that has led to greatly increased accuracy in predicting gene expression from DNA sequence.
— DeepMind (@DeepMind) October 4, 2021
Blog: https://t.co/21zIewaiKR
Paper: https://t.co/Bg1gPp4AWe 1/ pic.twitter.com/kxqgYcjfQ4
Additionally, the researchers have made their model public to advance the study of genes further. One can find the model here. DeepMind has recently made the source code for AlphaFold 2.0, helpful in predicting the shape of proteins, public as well.
What is Enformer?
DNA contains the genetic information that influences everything from eye colour to illness and disorder susceptibility. There are roughly 20,000 sections of DNA in the human body; we call them genes that contain instructions about the amino acid sequence of proteins. These genes perform various biochemical functions inside the cell. Despite this, these genes comprise less than 2% of the genome. The remaining base pairs in the genome are referred to as “non-coding,” and they include less well-understood instructions on when and where genes should be created or expressed in the human body. Nevertheless, they account for 98 per cent of the 3 billion “letters” in the genome.
The basic idea behind Enformer is to better understand variants in the non-coding genome and predict the effects of any variants on gene expression in both natural genetic and synthetic variants. Moreover, previous works on gene expression have used convolutional neural networks as fundamental building blocks; however, its inability to model the influence of distal enhancers on gene expression was a bottleneck for accuracy. Hence, the newly developed model is out for rescue.
The research has introduced a neural network architecture based on self-attention towards this goal. “We frame the machine learning problem as predicting thousands of epigenetic and transcriptional datasets in a multitask setting across long DNA sequences. Training on most of the human and mouse genomes and testing on held out sequences, we observed improved correlation between predictions and measured data relative to previous state-of-the-art models without self-attention,” as per the paper. Look at the figure to understand:
- Enformer is trained to predict human and mouse genomic tracks at 128-bp resolution from 200 kb of input DNA sequence.
- Enformer outperforms Basenji2 — state-of-the-art model, and
- Enformer consistently outperforms Basenji2 across all four assay types.
Image Credits: DeepMind paper
The major purpose of this new approach is to forecast which changes to the DNA letters, commonly known as genetic variants, would affect the gene’s expression. Enformer outperforms earlier models in predicting the impact of genetic variants on gene expression, both in natural genetic variants and synthetic variants that change critical regulatory sequences. This characteristic helps decipher the expanding number of disease-associated variations discovered in genome-wide association studies.
Tracing a bit of history
In 1990, an international scientific research project, i.e. Human Genome Project (HGP), saw its inception. The project’s goal was the complete mapping and understanding of all the genes (genome) of human beings. After almost 13 years, the mission to sequence three billion DNA letters in the human genome was completed in April 2003. The Human Genome Project’s completed sequence covers approximately 99 per cent of the human genome’s gene-containing regions and has been sequenced to a precision of 99.99 per cent. The achievements of the project over the years can be seen below.
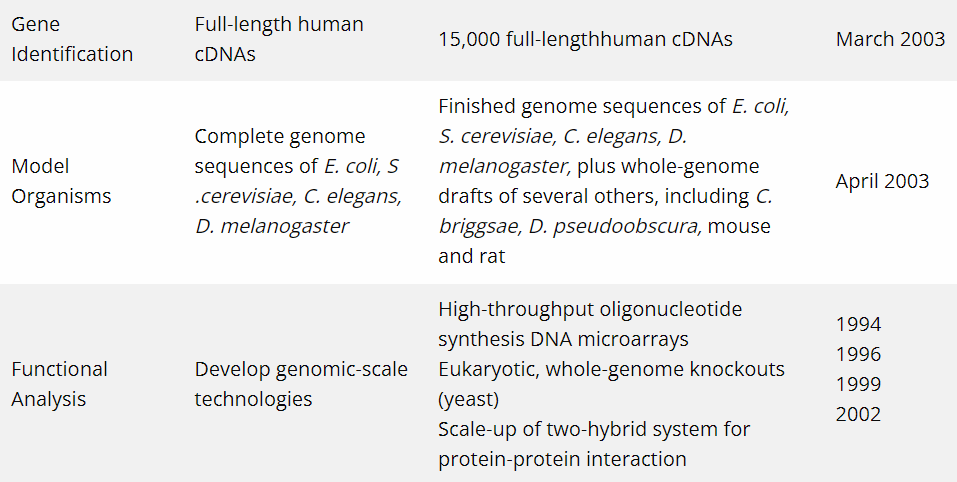


Image Credits: National Human Genome Research Institute
Inspired by HGP, in 2020, the Ministry of Science and Technology launched an ambitious gene mapping Genome India Project (GIP) in collaboration with 20 institutes, including IISc and IITs, for a period of three years. The intention is to build a grid of the Indian “reference genome” to identify and understand the type and nature of diseases and map the genetic diversity in India that will ultimately help in personalised medicine.
Enformer from DeepMind and various national and international projects are steps toward understanding the complexities of the genome sequence. Recent developments validate the fact that AI can play a much larger part when it comes to “genome” mapping. More such initiatives and research in this direction can further help in exploring new possibilities.