|
Listen to this story
|
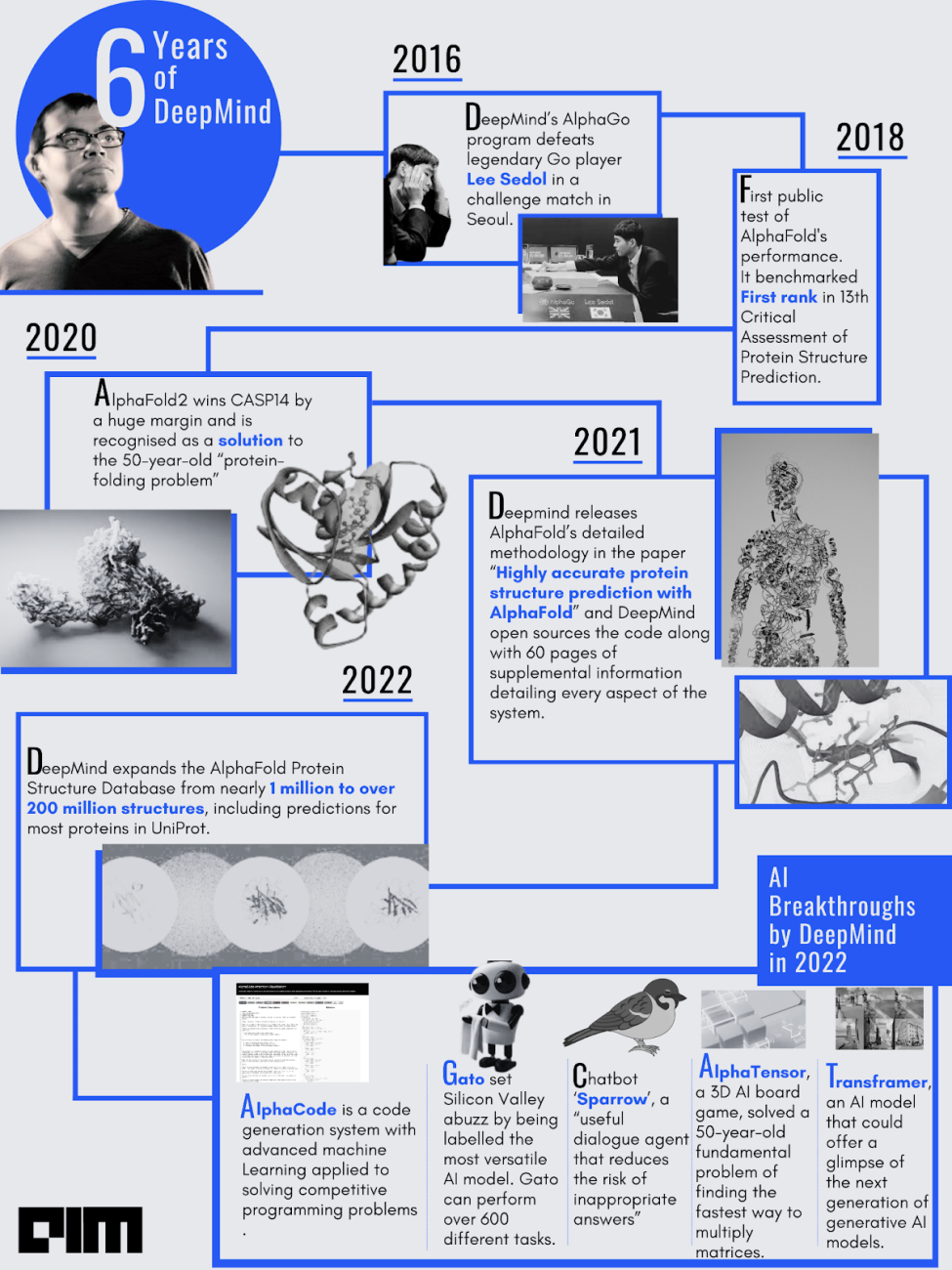
In July 2021, a London-based subsidiary of Alphabet, DeepMind, delivered the revolutionary answer to the decades-long ‘protein-folding problem’ in the history of AI research: AlphaFold. The open-source AlphaFold can accurately predict 3D models of protein structures from 1D amino acid sequences, which is accelerating scientific research in every field of biology and life science.
In an exclusive interaction with Analytics India Magazine, Pushmeet Kohli, head of research (AI for science, robustness and reliability) at DeepMind, shared the importance of AI for good, alongside his experience of being part of the revolutionary project and more.
“Science has given agency to humanity, and there are limits to our understanding of nature,” said Kohli, saying that the pandemic made it clear that we have no control over nature.
He further said that science has broadened our understanding of nature and has given us more means to leverage AI, which he believes will be one of the most powerful technologies that we as species can leverage to benefit science. “I think there is nothing more meaningful that one can do,” said Kohli along the lines of the importance of using AI for the greater good.
Protein Folding Models
“AlphaFold is a great example of how we can leverage AI because proteins are essentially the building blocks of all life. We are essentially a sort of big collection of proteins. It’s not just us; every single living thing on the planet is made up of these proteins,” said Kohli.
Further, he said that they did not completely understand what the structures are and what the function of all these proteins is. In that respect, Kohli believes that AlphaFold is a great watershed moment because it shows what AI can do in broadening the scientific community’s understanding of this important topic.
The function of a protein is directly related to its structure. For instance, like a key fitting into a lock, antibody proteins fold into forms that allow them to accurately detect and target particular foreign bacteria. Therefore, understanding how proteins will fold into shapes is crucial to understanding how organisms function and, eventually, how life itself works.
Merely 17% of the roughly 20,000 proteins in the human body had their 3-D structures known prior to AlphaFold. Enter AlphaFold and now 3-D structures for nearly the entire (98.5%) human proteome. This is a giant leap considering drug discovery is now easier.
AlphaFold
AlphaFold predicts protein structures through three distinct deep-learning neural network layers. It was trained on thousands of available proteins and their structures found in the Protein Data Bank (PDB). The first layer is made up of a variational autoencoder stacked with an attention model, which generates real-looking fragments based on a single sequence’s amino acids. The contact map, a 2D representation of amino acid residue distance, is projected onto a single dimension for input into the CNN (Convolutional neural network) in the first sublayer to optimise inter-residue distances. The second sublayer refines a scoring network, which measures how well the 3D CNN-generated substructures resemble proteins. After regularising it, they add a third neural network layer to compare the produced protein to the actual model.
Kohli said that his team’s target was to understand what the structure of proteins is, as every living creature’s tissue and cell have these proteins. AlphaFold predicts the structure of a protein. “But proteins are not always in a static state. They can be in multiple states according to their function or in the presence of other ligands. So, there remain many questions around how proteins interact with each other or with another set of ligands, and what energy they require to go from one state to another, among others, and our team is interested in working with them to find answers,” he added.
DeepMind used AlphaFold to predict the protein structures of the COVID-19 outbreak—SARS-CoV-2. Before making it public to the research community, the findings were reviewed by scientists at the Francis Crick Institute in the UK. The membrane protein, protein 3a, nsp2, nsp4, nsp6, and papain-like C-terminal domain are among these proteins. These protein structures were created to aid in the discovery of new medications and therapies in the fight against COVID-19 and may contain docking sites for those substances.