Google has proposed a new method, DeepNull, to model the complex relationship between covariate effects on phenotypes to improve Genome-wide association studies (GWAS). Google has released the DeepNull as open source software, along with a Colab notebook tutorial.
DeepNull models the nonlinear effect of covariates on phenotypes. It is simple to use and requires only a minimal change to existing GWAS pipeline implementations.
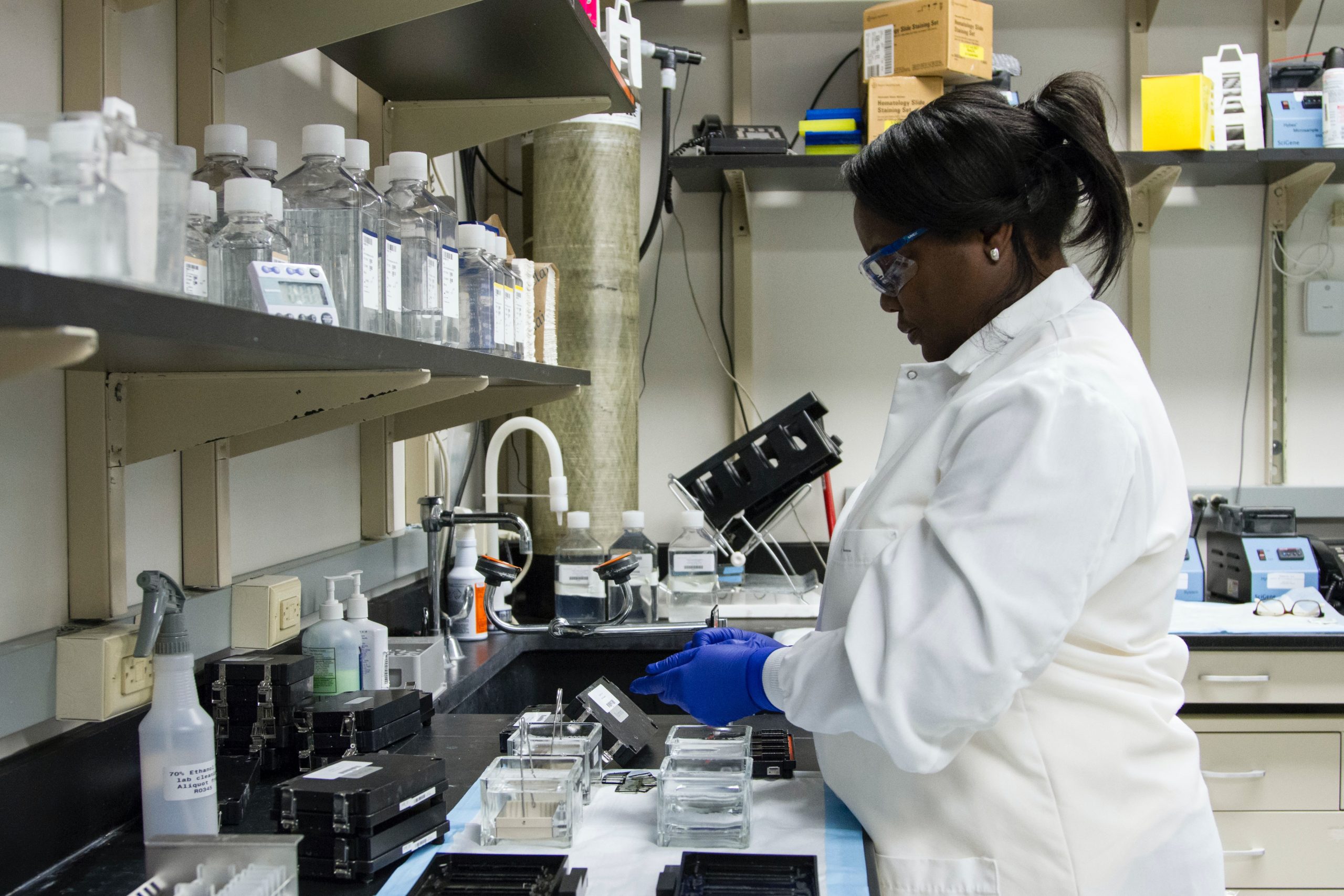
Genetic studies look for variants associated with different phenotypes (e.g., risk of diseases such as Glaucoma or observed phenotypic values such as high-density lipoprotein (HDL), low-density lipoproteins (LDL), height, etc).
GWAS are used to associate genetic variants with complex traits and diseases. The interactions between phenotypes (such as age and sex) and principal components (PCs) of genotypes, must be adjusted for as covariates to find out the association strength between genotype and phenotype. Covariate adjustment in GWAS can increase precision and correct for confounding. However, the assumption of linear and additive contributions of covariates doesn’t account for the underlying biology. To address this, Google researchers sought a method to more comprehensively model and adjust for the interactions between phenotypes for GWAS.
Working principle
DeepNull trains a deep neural network (DNN) to predict phenotype using all covariates in a 5-fold cross-validation. The prediction is an additional covariate in the association test. DeepNull is simple to use and requires only a minimal change to existing GWAS pipeline implementations.
DeepNull achieves results comparable to a standard GWAS when the effect of covariate on the phenotype is linear and can significantly outperform a standard GWAS when the covariate effects are nonlinear. DeepNull is open source and is available for download from GitHub or installation via PyPI.